Confirmatory Factor Analysis (CFA) in AMOS
Confirmatory Factor Analysis (CFA) in AMOS
Confirmatory Factor Analysis (CFA) is a method of Structural Equation Modeling (SEM) that is used to determine the observed variables’ contribution to the latent variables.
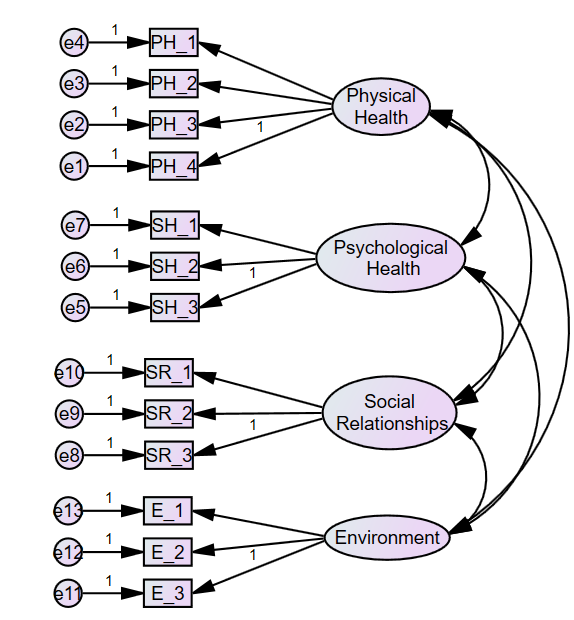
CFA is conducted for determining the variable of quality of life, which has four sub-constructs, namely, Physical Health, Psychological Health, Environment, and Social Relationships.
The following steps need to be followed to create CFA in AMOS:
Open IBM SPSS Amos and save the file by selecting File > Save. The following window will open.
Select the highlighted icon to load the data in AMOS.
‘Data Files’ dialog box will open. Select the data file by clicking ‘File Name’ option in the dialog box. Then click on ‘Ok’.
Draw the path diagram using the highlighted icon.
Click on the latent variable four times because physical health is affected by four factors. Four observed variables would be drawn.
The observed variables could be rotated using the highlighted icon.
Each variable needs to be specified using the imported dataset, for which the highlighted icon has to be selected.
‘Variables in Dataset’ dialog box will open.
Each variable has to be dragged from this dialog box to the drawn boxes of observed variables.
Double click on the latent variable and name it.
‘Object Properties’ dialog box will open.
In this case, it is named as ‘Physical health’.
Similarly, other latent variables need to be created. In this case, four latent variables have been created.
The unobserved variables must be named by clicking on ‘Plugins’ > ‘Name Unobserved Variables’
Add covariance to the latent variables using the highlighted icon.
Draw the covariance by connecting all latent variables.
Select the highlighted option to select the analysis properties.
Tick all the options mentioned in the figure under the ‘Output’ group.
Run the model using the highlighted icon “Calculate Estimate”.
Click on the highlighted icon ‘View Text’ to see the output.
Result file will be obtained.
Click on ‘Label’ in the ‘Options’ dialog box to see the name of variables in the result.
Estimates value for SH_1 is very less in the ‘Standardized Regression Weights’ table. Thus, that item will be removed and the model will be run again.
The following criteria need to be considered to check the model’s fitness.
CMIN/DF | Less than 5 |
NFI | Greater than 0.9 |
RFI | Greater than 0.9 |
IFI | Greater than 0.9 |
TLI | Greater than 0.9 |
CFI | Greater than 0.9 |
RMSEA | Less than 0.08 |
In this model, all the criteria were satisfied except CMIN/DF and RMSEA value.
This value could be improved by connecting error terms that have the highest ‘Modification Indices’ value.
Here, modification indices value for e2 <–> e4 is 60.896. Thus, e2 and e4 will be connected.
The model has to be run again.
Now, the CMIN/CF value would be less than 5.
Output of Confirmatory Factor Analysis
We will get ‘Regression Weights’ table. The values in the ‘Estimates’ column are values of unstandardized estimates.
It can be inferred from the table that all items were highly significant at p<0.01.
Then, we will get the table for Standardized Regression Weights. In this study, all items had standardized estimates, which were between 0.629 and 0.994, suggesting high loadings for quality of life.
It could be inferred that the model is a good fit, because CMIN/DF value is less than 0.5 in the CMIN table.
In the ‘Baseline Comparisons’ table, NFI (Normed Fit Index), RFI (Relative Fit Index), IFI (Incremental Fit Index), TLI (Tucker Lewis Index) and CFI (Comparative Fit Index) values were greater than 0.9, suggesting that the model is a good fit.
The RMSEA (Root Mean Square Error of Approximation) value of 0.078 is within the acceptable limit of <0.08 in the ‘RMSEA’ table.
Covariances were found to be highly significant among all latent constructs with estimates ranging between -0.101 to 0.632 for the relationships.
Significant correlations were found among all latent constructs.
Data: CFA_Data_1.sav